what human cell lines are aeasy to transfect
Abstract
With the rise of new powerful genome applied science technologies, such as CRISPR/Cas9, cell models can be engineered effectively to accelerate bones and illness research. The most critical step in this procedure is the efficient delivery of foreign nucleic acids into cells by cellular transfection. Since the vectors encoding the components necessary for CRISPR/Cas genome applied science are ever large (nine–nineteen kb), they event in low transfection efficiency and prison cell viability, and thus subsequent selection or purification of positive cells is required. To overcome those obstacles, we here show a non-toxic and non-viral delivery method that increases transfection efficiency (upwards to xl-fold) and jail cell viability (upwardly to 6-fold) in a number of difficult-to-transfect human cancer cell lines and primary blood cells. At its core, the technique is based on calculation exogenous small plasmids of a defined size to the transfection mixture.
Introduction
CRISPR/Cas has revolutionized genome engineering science of biological systems due to its easy design, target site specificity, and scalability for high-throughput applications. It allows gene deletions, enhancing or inhibiting gene expression in vitro and in vivo. The components of the CRISPR/Cas organization (including guide RNAs) are often encoded on large extrachromosomal expression vectors (9–19 kb) that are delivered into cells via transfection. Due to the big size, these vectors are notoriously hard to transfect and cause high cell death, which prohibit downstream analysesi,2.
Cell transfection method development has resulted in safer viral vectors (biological)3, new polymers and lipids (chemical)4, and particle delivery devices (physical)4. Viral-mediated delivery (transduction) leads to the highest efficiencies only requires higher biosafety level laboratory settings and ethical approving when used in enquiry or in the clinic5. We overcame limitations of current electroporation-based transfections by adding appropriate amounts of small (~3 kb) to large (ix–15 kb) vectors, which increased transfection efficiency and prison cell viability. Due to its easy implementation in current transfection protocols, this strategy may be broadly applicative in basic and applied research.
Results
Pocket-sized vectors ameliorate transfection efficiencies
Standard transfection via electroporation (Fig. 1a, Table 1) of a fifteen kb CRISPR-GFP vector into hard-to-transfect human lung cancer cells (A549) showed extremely depression transfection efficiency (4.2%) and high prison cell expiry (91%) (Fig. 1b, c). In dissimilarity, co-transfection of equal mass of a small empty vector (3 kb) together with the large CRISPR-GFP vector (xv kb) drastically increased transfection efficiency (twoscore%) and reduced cell death (45%) (Fig. 1b, c).
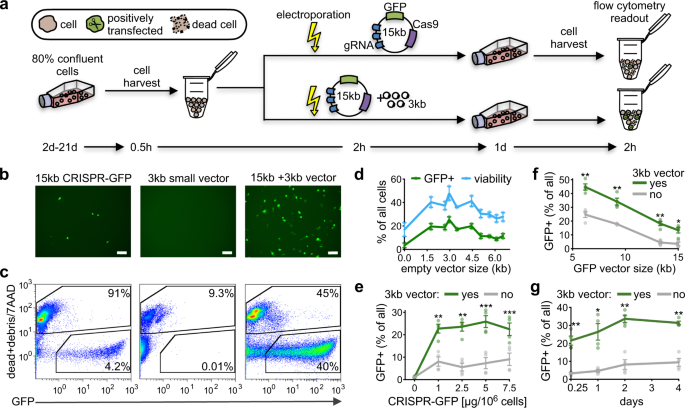
a Schematic overview of the cell transfection setting. Electroporation-mediated transfection (lightning bolt) of a big CRISPR-GFP vector (xv kb) without (above) and with (beneath) a minor vector (three kb). Duration in days (d) and hours (h) for each experimental procedure is indicated. b, c Microscopy images and flow cytometry plots (gating of GFP+ and 7AAD dead-cell mark) of hard-to-transfect A549 cells 24 h after electroporation (left: fifteen kb CRISPR-GFP vector lone, center: iii kb modest vector alone, correct: co-transfection of 15 kb CRISPR-GFP and 3 kb small vector). Scale bar: 100 µm. Amounts of vector and electroporation conditions tin can be found in Table i. d Line graph illustrates percent transfection efficiency (greenish) and cell viability (bluish) upon co-transfection of a big 15 kb vector with minor vectors of varying sizes (1.8–6.5 kb) in A549 and MCF7 cells (northward = 4, mean ± SEM). due east Line graph demonstrate the percent transfection efficiency later on co-transfection of a large CRISPR-GFP vector (15 kb) of varying concentrations without (gray) and with (dark-green) a modest vector (three kb) in A549 and MCF7 cells (n = 4, hateful ± SEM). f Line graph demonstrate the percentage of viable GFP + cells subsequently co-transfection of large GFP vectors (6.5–15 kb) without (gray) and with (green) a small vector (3 kb) in A549 and MCF7 cells (northward = 4, mean ± SEM). g Line graph demonstrate the percentage transfection efficiency after co-transfection of a big CRISPR-GFP vector (15 kb) without (grey) and with (green) a minor vector (iii kb) in A549 and MCF7 cells from half-dozen h (0.25d) to 4d after transfection (north = 4, mean ± SEM). Statistics: paired ii-tailed t-test, *p < 0.05, **p < 0.01, ***p < 0.001.
We next tested if the size of the pocket-size vector influences transfection efficiencies by using a range of small vectors (1.viii–6.5 kb) (Fig. 1d, Supplementary Fig. 1, Table 2). On boilerplate, co-transfection of the large CRISPR-GFP vector with small vectors increased transfection efficiency past 12.2% (four.9-fold change) and prison cell viability by xvi.6% (1.ix-fold change). Of all vectors tested, the small vector of 3 kb showed the highest increment in transfection efficiency (average of 21.4% or 6.eight-fold increase) (Fig. 1d, Supplementary Fig. 1f–h). Furthermore, transfection of solely modest vectors did not significantly alter prison cell viability when compared to mock transfection (paired two-tailed t-test, p-value > 0.05) (Supplementary Fig. 2). Since all pocket-size vectors improved transfection efficiencies, we speculate that our approach has been employed unknowingly when co-transfecting gRNAs and CRISPR/Cas components on separate vectors6. Transfection of cells with increasing amounts of CRISPR-GFP vector (15 kb, 0–7.v µg) did not result in an increment of GFP+ cells but rather lead to a higher number of dead cells (Fig. 1e, Supplementary Fig. 3). Co-transfection of a stock-still corporeality of the small vector (3 kb, five µg) increased the number of GFP+ cells consistently (iv.3-fold change on boilerplate) and increased the number of feasible cells (i.9-fold change on average) (Fig. 1e). This suggests that the size just non the amount of the large vector affect transfection efficiencies.
To validate whether the increase in transfection efficiency and jail cell viability was dependent on the size of big CRISPR vectors, we electroporated cells with a range of different GFP vectors (six.5–15 kb). We found a gradual decrease in transfection efficiency (from ca. 25 to 4%) and jail cell viability (from ca. 36 to 15%) with increasing GFP vector size (Fig. 1f, Supplementary Fig. 4). This vector size-dependency on transfection efficiency tin exist considered when designing futurity CRISPR vectors. Remarkably, transfection efficiency and cell viability improved considerably upon co-transfection of the small vector (3 kb) with all tested GFP vectors. Overall, we found an average increase in transfection efficiency of 15% (range: 6–25%) irrespective of the size of the GFP vector. Additionally, GFP expression in cells that were transfected with a large CRISPR-GFP vector (15 kb) in the presence of small (three kb) plasmids showed a stable enhancement in transfection efficiencies lasting for several days (6 h to 4d after transfection) (Fig. 1g, Supplementary Fig. 5).
Minor vectors increment transfection efficiency in numerous cell types
Patient-derived homo cell lines have been intensively used in research to investigate molecular mechanisms explaining diseases also as to identify and test pharmaceutical compounds for therapeutic purposes nether well-defined and reproducible conditions. Furthermore, main cells from the peripheral blood isolated from patients can exist genetically modified and used every bit immunotherapy in clinical applications. In lodge to determine whether our approach tin be employed in other intensively studied, difficult-to-transfect or primary prison cell types, nosotros co-transfected small (3 kb) and large (15 kb) vectors to measure transfection efficiencies and cell survival in various adherent and non-adherent cancer cell lines (Huh7 and HepG2 (liver), PC3 (prostate), MCF7 (breast), HEK293 (kidney), A549 (lung), SH-SY5Y (neuronal), HL-60 (leukaemia)) as well as peripheral blood mononuclear cells (PBMCs) and purified CD8 T cells (Fig. 2a). In line with our previous observations, we confirmed a consistent improvement of transfection efficiency (upwardly to 36%) and cell viability (up to 46%) in all tested prison cell lines (Fig. 2b–e, Supplementary Fig. 6). In contrast to adherent cells, cells in pause had a very low transfection efficiency, which could probably be improved upon further optimization of the electroporation settings. However, co-transfection with the minor vector did even so ameliorate the number of GFP+ cells (Fig. 2b, d, Supplementary Fig. 6) but did not increase cell viability (Fig. 2c, e). Overall, we noticed that the increased percent of positively transfected cells correlated highly with the percentage of viable cells (Fig. 2f), suggesting that the manner of action of the small vector is to better viability of positively transfected cells. To better understand the underlying mechanism, we examined dependencies of transfection efficiencies on the small vector conformation (Supplementary Fig. 7a–e). Linearizing the minor vector did not pb to any difference in transfection efficiency when compared to its circular version, suggesting that round and linearized plasmids are equally capable of enhancing the transfection efficiency. Nosotros also considered vector sequence content and features and establish that neither GC content nor specific motifs in the encoded Deoxyribonucleic acid of the small vector improved transfection efficiencies (Supplementary Fig. 7f).
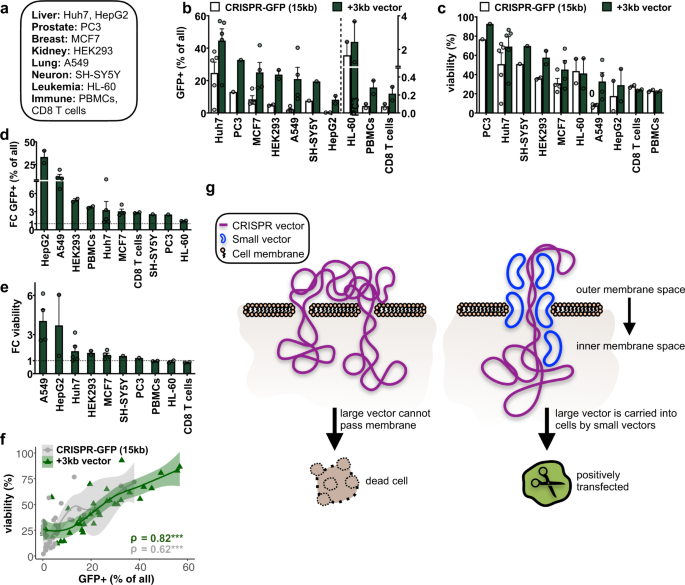
a Diverse human cancer cell lines and freshly isolated principal allowed cells were electroporated. b, c Bar graphs depicting the percentage increase of b transfection efficiency and c cell viability upon co-transfection of the CRISPR-GFP (15 kb) vector without (white) or with (night green) a modest vector (3 kb) in the tested adherent (left of the dotted line) and non-adherent (correct of the dotted line) cells (n = i–6, mean + SEM). Jail cell types are ranked by decreasing transfection efficiency or cell viability later on adding the small vector. Amounts of vector and electroporation conditions can be plant in Table ane. d, e Bar graphs illustrate the fold change (FC) in the number of d GFP+ and e viable cells after adding a small 3 kb vector ranked by FC-enrichment in the tested cells. f Plot shows Spearman's rank correlation coefficients (ρ) and p-values (***p < 0.001) of transfection efficiencies (x-axis) and cell viability (y-centrality) (in percent) without (gray) or with (light-green) co-transfection of a small vector (3 kb) (north = 26, 95% conviction interval). thousand Model explaining increased transfection efficiencies of large CRISPR vectors (royal) upon addition of small-scale vectors (blue). Membranes and nuclear pores are coated by modest vectors and thereby facilitates efficient commitment into the nucleus and subsequent molecular action of the large vector.
Finally, we tested our approach in chemic transfection methods. We employed the commonly used Lipofectamine 3000 and generated liposomes containing the large CRISPR-GFP vector (xv kb) in the presence and absence of a small vector (iii kb). Like to the results obtained subsequently electroporation, the add-on of the small vector enhanced transfection efficiencies in all of the tested cell types (Supplementary Fig. eight). However, in contrast to electroporation-based transfection, inclusion of the small vector in the liposomes slightly decreased the viability (Supplementary Fig. 8).
Give-and-take
We postulate that minor vectors can quickly pass cell and nuclear membranes and large vectors can move on this flow of small vectors into the cell. By co-travelling with small-scale vectors, larger vectors may enter the cell without getting entangled in two or more open membrane pores, which ensures proper plasmid uptake and membrane reclosure thereby preventing cell deceasevii (Fig. 2g). This model would explain the results we take obtained using electroporation of adherent cells, but it explains neither the poor viability of suspension cells nor why the small vector improves liposome transfection efficiency. Compared to the adherent cells, cells in break are generally smaller and have non undergone trypsinization. Trypsinization cleaves the bonds that stretch the membrane of the adherent cells and may thus upshot in a less compact and a more relaxed membrane construction. This may facilitate the germination of larger membrane pores after electroporation, assuasive better accessibility of pocket-sized plasmids and thereby smoother membrane passage of larger vectors (Fig. 2g). The deviation in cell death later on co-transfection with modest vectors may be further explained by different responses of DNA sensors triggering programmed cell expiry8. Dna sensors might be inert in response to small plasmids merely highly active when sensing large plasmids. Withal, this phenomenon is speculative, and more biophysical work is required to resolve the exact underlying mechanism.
An culling arroyo to our strategy has previously been publishedii,9. Different transcribing the CRISPR/Cas9 organisation inside the host prison cell as we propose, a recombinant CRISPR/Cas9 ribonucleoprotein tin can be formed in vitro prior to electroporation. This arroyo yielded higher transfection efficiencies in principal claret cells than we report. However, a recombinant poly peptide has to be produced and the ribonucleoprotein complexes must be electroporated immediately after formation. Compared to this strategy, our approach of adding a small vector to the transfection mixture is simpler, cheaper and less time-consuming.
In summary, we discovered that electroporation and lipofectamine-based cell transfection of cancer cell lines and master cell types can be improved by adding a pocket-sized vector to the transfection mixture. Equally CRISPR technology is universally applicable and will continue to develop further, our optimized, simple and non-hazardous transfection approach will have numerous applications in clinical biomedicine and industrial biotechnology.
Methods
Cell culture and cell lines
HepG2, Huh7, PC3, SH-SY5Y, HEK293, MCF7, HL-60, and A549 cancer jail cell lines were obtained from American Type Culture Collection (ATCC). All cell lines were mycoplasma-gratis when periodically tested with Mycoplasmacheck (Eurofins Genomics) or MycoProbe (R&D). Cells were cultured in T-75 flasks at 37 °C and 5% CO2 temper using medium supplemented with 1/100 Penicillin/Streptomycin (Sigma) and 10% fetal bovine serum (Hyclone). Huh7, HepG2, A549, HEK293, and MCF7 were cultured in Dulbecco's Modified Hawkeye Medium (DMEM, Sigma), HL-sixty was cultured in Roswell Park Memorial Institute (RPMI) 1640 (Sigma), and PC3 and SH-SY-5Y were cultured in DMEM:F-12 (i:one) medium (Gibco). To ensure authenticity, jail cell lines were initially genotyped by short-tandem repeat genetic profiling (STR) using the PowerPlex_16HS_Cell Line panel and analyzed using Practical Biosystems Gene Mapper ID v3.2.i software by the external provider Genetica Dna Laboratories (LabCorp Specialty Testing Group) and continuously assessed phenotypically. Cells were split at ~70–ninety% confluency past aspirating the medium, gently washing with phosphate buffered saline (PBS, Sigma) and detaching them with iii mL of a trypsin-EDTA solution (Sigma) for three–5 min. Trypsin was inactivated with a minimum of 10-fold surplus of civilisation medium before a cell fraction was passaged.
Vectors
pLV hU6-sgRNA hUbC-dCas9-KRAB-T2a-GFP10 (15.0 kb), FC3-GFP (half-dozen.two kb), SpCas9(BB)-2A-GFP-prickle (9.2 kb), pCAGGs-jmj1dc-IRES-GFP (13.three kb), pUC19-no-LacZ (one.8 bp), pUC19 (two.seven kb), pBlueScript (3.0 kb), pH6HTC (3.5 kb), pH6HTC-STMN (4.4 kb), pH6HTC-PKM (5.1 kb), pH6HTC-CCT (five.six kb), pH6HTC-CTCF (6.ii kb), pH6HTC-D9 (6.5 kb).
Electroporation
Cells were electroporated using the NEON electroporation system (Invitrogen). Briefly, cells grown to 70–xc% confluency were harvested and pelleted at 500 g for 5 min at room temperature. Cells were resuspended in PBS, counted, and spun down at 500 one thousand for five min at room temperature. The appropriate amount of plasmid DNA (see Tabular array ane) were transferred into a sterile 1.5 mL microcentrifuge tube. After aspirating PBS from the prison cell pellet, the cells were resuspended in Resuspension Buffer R to 1.0 × 107 cells/mL. Cells were gently mixed to obtain a single prison cell suspension and added to the tube containing plasmid Dna. The cells were mixed gently with the plasmids without creating any air bubbling. To avert unnecessary prison cell death, the electroporated cells were straight plated into a pre-heated phenol red-costless medium without any antibiotics. Electroporation settings used are found in Tabular array 2.
Liposomal transfection
Liposomal transfection was done with lipofectamine 3000 according to the manufacturer'south instructions. All cells were plated in 24-well plates (110,000/well HepG2 and MCF7, lxx,000/well Huh7, and fifty,000/well A549). Mix one: 25 μL Opti-MEM (Gibco) + ane.5 μL Lipofectamine 3000. Mix 2: 25 μL Opti-MEM + 250 ng pBluescript + 250 ng GFP-vector + 1 μL P3000 reagent.
Transfection efficiency and cell viability measures
Transfection efficiency was measured later 24 h (unless stated differently) by period cytometry (FACSNavios, Beckman Coulter, Navios Cytometry Listing Mode Data Acquisition and Assay Software version 1.3) by gating cells for GFP and 7AAD (FlowJo version 8.two, Supplementary Fig. ane). In order to collect all potentially dead cells, both the supernatants and the adherent cells (harvested by trypsinization) were collected. Cells were washed in 1xPBS with 1% BSA (Sigma), followed by staining in 100 μL buffer with v μL 7AAD viability staining solution (eBioscience) for 15 min on ice in the night. Cells were acquired directly without washing away the staining buffer by flow cytometry. In some experiments, microscopy images were taken using a Zoe Fluorescent Prison cell Imager (Bio-Rad, software version 002.257.011215).
Chief immune cell isolation
Buffy coats were obtained from bearding healthy volunteers after informed consent and according to institutional guidelines (Karolinska Academy Hospital, Stockholm, Sweden). PBMCs were isolated using Ficoll-Paque (GE Healthcare) as previously described11. CD8 T cells were isolated using magnetic associated cell sorting (Miltenyi) according to the manufacturer'south instructions. PBMCs and CD8 T cells were rested overnight at 37 °C and 5% CO2 temper in RPMI supplemented with 1/100 Penicillin/Streptomycin and ten% fetal bovine serum prior to electroporation.
Statistics and reproducibility
The data were plotted and analysed with GraphPad Prism (version viii.4.2), Microsoft Excel (version sixteen.37), R (version three.6.1), and ggplot2 (version three.3.0). Replicates are divers as private passages of cancer cell lines or as individual donors for primary immune cells. Data are represented as mean + or ± SEM and sample sizes are presented in each effigy fable. Statistical significance of differences among groups was determined by paired two-tailed student's t-examination. A p-value smaller than 0.05 (p < 0.05) was considered to be statistically meaning.
Reporting summary
Further information on inquiry design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All data generated or analyzed during this study are included in the article. Raw data for graphs tin be institute in Supplementary Information 1. Additionally, all relevant data are bachelor from the authors upon request.
References
-
Lesueur, L. Fifty., Mir, L. M. & André, F. M. Overcoming the specific toxicity of big plasmids electrotransfer in primary cells in vitro. Mol. Ther. Nucleic Acids 5, e291 (2016).
-
Roth, T. L. et al. Reprogramming human T jail cell function and specificity with non-viral genome targeting. Nature 559, 405–409 (2018).
-
Münch, R. C. et al. Displaying high-affinity ligands on adeno-associated viral vectors enables tumor prison cell-specific and safe gene transfer. Mol. Ther. 21, 109–118 (2013).
-
Colina, A. B., Chen, Grand., Chen, C. Yard., Pfeifer, B. A. & Jones, C. H. Overcoming cistron-delivery hurdles: physiological considerations for nonviral vectors. Trends Biotechnol. 34, 91–105 (2016).
-
Schlimgen, R. et al. Risks associated with lentiviral vector exposures and prevention strategies. J. Occup. Environ. Med. 58, 1159–1166 (2016).
-
Mali, P. et al. RNA-guided homo genome engineering science via Cas9. Science 339, 823–826 (2013).
-
Stewart, K. P., Langer, R. & Jensen, K. F. Intracellular delivery past membrane disruption: mechanisms, strategies, and concepts. Chem. Rev. 118, 7409–7531 (2018).
-
Maelfait, J., Liverpool, Fifty. & Rehwinkel, J. Nucleic acid sensors and programmed prison cell death. J. Mol. Biol. 432, 552–568 (2020).
-
Schumann, K. et al. Generation of knock-in primary human T cells using Cas9 ribonucleoproteins. Proc. Natl Acad. Sci. The states 112, 10437–10442 (2015).
-
Thakore, P. I. et al. Highly specific epigenome editing by CRISPR-Cas9 repressors for silencing of distal regulatory elements. Nat. Methods 12, 1143–1149 (2015).
-
Søndergaard, J. Northward. et a. Dendritic cells actively limit interleukin-10 production nether inflammatory atmospheric condition via DC-SCRIPT and dual-specificity phosphatase 4. Front. Immunol. nine, 1–thirteen (2018).
Acknowledgements
Nosotros would like to thank the KI Innovation office, specifically Åsa Kallas and Marker Didmon for helpful discussions. This work was supported by the Knut & Alice Wallenberg foundation (KAW 2016.0174, C.One thousand.), Ruth & Richard Julin foundation (2017–00358 and 2018–00328, CK); SFO-SciLifeLab fellowship (SFO_004, C.K.), Swedish Research Quango (2019–05165), Chinese Scholarship Council (KG, C.K.), KI-Kid funding (2016–00189 and 2018–00904, C.K.), and the Nilsson-Ehle Endowments (J.N.South.). Open up access funding provided by Karolinska Institute.
Author information
Affiliations
Contributions
J.Due north.S., X.Y., and C.Thousand. conceptualized the project. J.N.S, K.G., C.S., and I.A. performed the laboratory experiments. J.N.S. did the analysis and visualized the data. J.N.S. and C.1000. acquired the funding. J.N.Southward. and C.Grand. wrote the original typhoon. All authors contributed to the review and editing process.
Respective writer
Ethics declarations
Competing interests
The authors declare no competing non-financial interests but the following potential competing fiscal involvement: a patent has been filed for the use of small vectors to increase cellular uptake of big vectors upon transfection. Bidder: Biotech & Biomedicine (Shenyang) Group Ltd. Authors: Claudia Kutter, Jonas Nørskov Søndergaard, and Xiushan Yin. Application number: 202010315646.4. Status of awarding: submitted, commencement example. Submitted to: National Intellectual Property Assistants, People'south Republic of China. Also from this patent application, the authors declare no additional competing financial interests.
Additional information
Publisher's note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed nether a Artistic Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, equally long as you give appropriate credit to the original author(south) and the source, provide a link to the Artistic Commons license, and bespeak if changes were made. The images or other third political party textile in this article are included in the commodity'due south Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the commodity'south Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you lot will need to obtain permission direct from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/past/4.0/.
Reprints and Permissions
Nigh this article
Cite this article
Søndergaard, J.N., Geng, Chiliad., Sommerauer, C. et al. Successful delivery of large-size CRISPR/Cas9 vectors in hard-to-transfect human cells using small plasmids. Commun Biol 3, 319 (2020). https://doi.org/10.1038/s42003-020-1045-7
-
Received:
-
Accepted:
-
Published:
-
DOI : https://doi.org/10.1038/s42003-020-1045-vii
Comments
By submitting a comment you agree to bide by our Terms and Community Guidelines. If yous find something abusive or that does not comply with our terms or guidelines delight flag it as inappropriate.
barnhartfashe1978.blogspot.com
Source: https://www.nature.com/articles/s42003-020-1045-7
0 Response to "what human cell lines are aeasy to transfect"
Post a Comment